Newly discovered plant enzymes could lead to more efficient -- and less costly -- ethanol production from cellulose
By Krishna Ramanujan
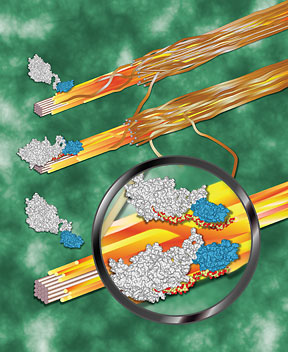
In a breakthrough that could make the production of cellulosic ethanol less expensive, Cornell researchers have discovered a class of plant enzymes that potentially could allow plant materials used to make ethanol to be broken down more efficiently than is possible using current technologies.
There is a growing recognition that corn ethanol is unlikely to provide a long-term solution, or one that is environmentally sustainable, and so scientists are turning to cellulose as an alternative.
Production of ethanol from cellulose in mass quantities that are priced competitively with corn-based ethanol has not yet been possible. And without the cellulosic ethanol, the national goal for ethanol production to reduce oil imports will be impossible to reach, experts say.
A critical step in producing cellulosic ethanol involves breaking down a plant's cell wall material and fermenting the sugars that are released. Current technologies use microbial enzymes called "cellulases" to digest the cellulose in grasses and such rapidly growing trees as poplars. The microbial enzymes have a structure that makes them very efficient at binding to and digesting plant cell wall material called lignocellulose (a combination of lignin and cellulose).
But now, a new class of plant enzymes with a similar structure has been discovered, potentially offering researchers new properties for producing ethanol even more efficiently.
"The bottleneck for conversion of lignocellulose into ethanol is efficient cellulose degradation," said Jocelyn Rose, Cornell assistant professor of plant biology. "The discovery of these enzymes suggests there might be sets of new plant enzymes to improve the efficiency of cellulose degradation."
The paper appears in the April 20 issue of the Journal of Biological Chemistry. Breeanna Urbanowicz, a graduate student in Rose's laboratory, was the paper's lead author.
For an enzyme to break down cellulose, a structure called a cellulose-binding module attaches to the cellulose. Once attached, a catalyst then breaks the cell wall material into small units, which can then be turned into ethanol. While researchers have known that plants have cellulaselike enzymes, it was previously thought that they did not have a cellulose-binding module, and so could not attach to cellulose or digest it very effectively -- until now.
"This is the first example of a cellulose-binding domain in a plant cell wall enzyme," said Rose.
While the new enzyme was found in a tomato plant, Rose and colleagues have evidence of a set of such plant proteins in many species that potentially could be used for biofuel production. Biofuel research may also help uncover exciting new uses for these enzymes, said Rose. Researchers may, for example, breed for plants with high levels of these proteins.
Though the scientists stress that more study is needed to understand how plants use this class of enzymes, Rose speculates that they may be needed when growing tissues rapidly expand and require loosening of tightly bound strands of cellulose, called microfibrils, that make up a cell wall's structure. The binding enzymes may also be part of the process of breaking down tissues, e.g., when fruits -- such as tomatoes -- soften.
Among others, co-authors included Carmen Catalá, a research associate previously working in the Department of Plant Biology, who originally identified the gene for the tomato enzyme, and David Wilson, Cornell professor of molecular biology and genetics.
Media Contact
Get Cornell news delivered right to your inbox.
Subscribe