New strategy IDs infertility-causing genes
By Krishna Ramanujan
Infertility affects roughly 15 percent of couples – with many cases resulting from faulty genetics – but the details of underlying genetic causes are largely unknown.
Now, Cornell researchers have developed an experimental strategy to identify infertility-causing mutations found in human populations. These mutations are known as single nucleotide polymorphisms, or SNPs, and are the most common type of genetic variation among people. Each SNP represents a difference in a single DNA building block, called a nucleotide.
“If we figure out whether a SNP is truly deleterious, then in the future when patients come in they can have their genomes sequenced to determine which SNPs they have. If we know which variation is good or bad, doctors will be able make a genetic diagnosis,” said John Schimenti, director of the Center for Vertebrate Genomics at Cornell and the paper’s senior author. Priti Singh, a postdoctoral fellow in Schimenti’s lab, is the first author of the study published Aug. 3 in the Proceedings of the National Academy of Sciences.
The standard way to identify disease-causing SNPs involves comparing the genomes of healthy and affected people to narrow down their chromosomal locations, then using algorithms to predict which SNPs are harmful. But because fertility is such a complex process involving many genes, this method has not worked.
So Schimenti and Singh developed a new strategy: They took a list of all the known infertility genes in mice – which have been well-established through experimentation that cannot be done with humans – then computationally identified the equivalent SNPs in humans through databases of human genetic variation.
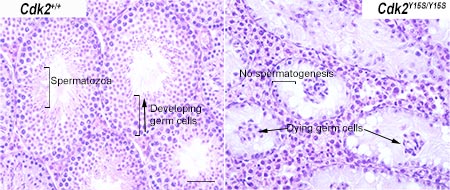
With this new list, the researchers explored SNPs of four essential genes that were predicted to cause infertility in humans. Using a genome editing technology, called CRISPR/Cas, Schimenti and Singh edited the homologous genes in mice so they mimicked the suspected human infertility SNPs. By growing the mice with these “humanized” infertility SNPs, they functionally tested if the human mutations caused infertility in the mice. Of the four SNPs tested, one caused infertility.
“We were surprised at how poor the existing, commonly-used algorithms were at predicting deleterious SNPs,” Schimenti said. “But with this technology, we are starting to identify those SNPs that really do cause infertility” in humans.
He predicts that in the future such advances will lead to personalized medicine where a person may have his or her genome sequenced, perhaps at birth, and doctors and patients will be able to determine the genetic health issues of every person.
“Right now, there is a self-imposed moratorium on genome editing in humans, but in the future, steps could be taken to repair the mutations,” Schimenti said.
Also, “a lot of people want to know why they are infertile” and whether the cause is genetic or environmental, he added, and this research is a first step toward such a diagnosis.
Schimenti has now teamed with Haiyuan Yu, assistant professor of biological statistics and computational biology at the Weill Institute for Cell and Molecular Biology, and has received a National Institutes of Health grant to use computational methods with genome editing in mice to identify more potentially infertility-causing SNPs.
The study was funded by the National Institutes of Health and the New York State Stem Cell Program.
Media Contact
Get Cornell news delivered right to your inbox.
Subscribe