New method of bacterial cell engineering can produce better, cheaper drug therapies
By Anne Ju
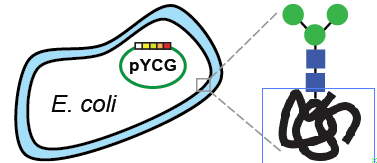
Therapeutic proteins, which provide cutting-edge treatments of cancer, diabetes and countless other diseases, are among today's most widely consumed biopharmaceuticals. By introducing bottom-up carbohydrate engineering into common bacterial cells, Cornell researchers have discovered a way to make these drugs cheaper and safer.
A research team led by Matthew DeLisa, associate professor of chemical and biomolecular engineering, has invented a novel method for engineering human therapeutic glycoproteins simply and quickly using E. coli bacteria as a platform. Their work is detailed online March 25 in Nature Chemical Biology.
Their methods are now being commercialized through a startup company, Glycobia Inc., which recently took up residence in Cornell's McGovern Family Center for Venture Development in the Life Sciences.
Glycoproteins are proteins that are modified at specific amino acid "acceptor" sites with complex carbohydrate structures, or oligosaccharides -- a basic human chemical reaction that's essential to life. That's why specifically designed, genetically engineered glycoproteins are so commonly used as drugs -- they bind to certain protein receptor sites and, for example, block cancer cells from multiplying. Among glycoproteins used to treat diseases today are monoclonal antibodies and interferons.
Current manufacturing methods rely mainly on costly, time-consuming mammalian culture cells, such as the Chinese Hamster Ovary (CHO) cell line. The process is also susceptible to viral contamination, further driving up production cost; in fact, in 2009 the biopharmaceutical company Genzyme temporarily shut down its Allston, Mass., plant after such a contamination occurred.
The Cornell research uses a bottom-up engineering method to assemble a synthetic pathway for the simple and quick production of a glycoprotein that forms the basis of many of today's therapeutic protein drugs, including, for example, the protein GCase, used in a drug that treats Gaucher's disease. To do so, they artificially introduced the machinery of glycosylation -- the chemical process by which proteins become glycoproteins -- into E. coli cells, rather than animal cells.
The synthetic pathway they designed, which can be tailored to many amino acid acceptor sites to make different drugs, starts with native enzymes in E. coli. Added to that was a mixture of four enzymes taken from yeast cells, which triggered the biosynthesis of a specific glycan (sugar structure) that resembles the core structure found in virtually all eukaryotic glycans. A fifth enzyme from the bacterium, Campylobacter jejuni, transferred these core glycans to pre-engineered protein acceptor sites, resulting in the desired glycoproteins.
The researchers are now working to improve their approach that they call "glycans by design" -- using the enzyme-based protein production method to specifically tailor sugar structures to make many different glycans and glycoproteins.
The research was supported by National Science Foundation; New York State Office of Science, Technology and Academic Research; and the National Institutes of Health.
Media Contact
Get Cornell news delivered right to your inbox.
Subscribe