Scientists unwind mystery behind DNA replication
By Linda B. Glaser
The molecules of life are twisted. But how those familiar strands in DNA’s double helix manage to replicate without being tangled up has been hard to decipher. A new perspective from Cornell physicists is helping unravel the mystery.
Researchers approached the problem from a topological perspective, asking what impact the helix shape itself has on DNA replication. Using eukaryotes – which comprise the vast majority of living things – as their model system, they found that the intrinsic mechanical properties of the chromatin (a complex of DNA and proteins) determine how the chromatin fibers will entwine.
This topology is crucial to the successful separation of newly replicated DNA: If the fibers twist too tightly too early, then the molecules are unable to properly segregate during cell division.
“This research highlights the importance of physical principles in fundamental biological processes,” said lead author Michelle Wang, professor of physics in the College of Arts and Sciences and a Howard Hughes Medical Institute Investigator.
The paper, “Synergistic Coordination of Chromatin Torsional Mechanics and Topoisomerase Activity,” published Oct. 17 in Cell.
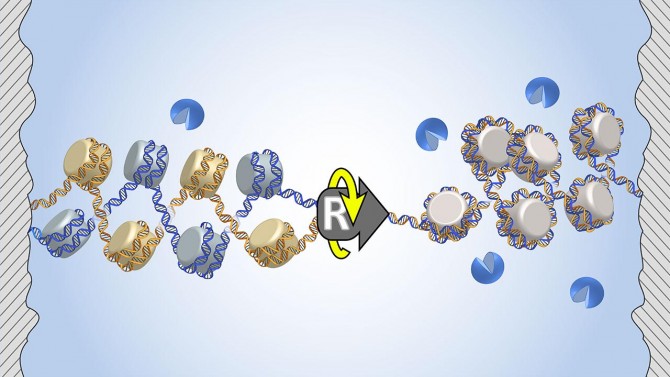
During DNA replication – as a replisome splits two DNA strands and moves forward – the DNA must also twist around the helical axis. This puts DNA under a great deal of torsional stress, which then results in an extra twist in the DNA.
The question: Where does the extra twist go? If the extra twist goes only to the front of the replisome, then the two daughter DNA molecules would not get tangled up, so they can separate. However, if the extra twist goes to the back of the replisome, then the two daughter DNA molecules would get tangled up and could not separate. This would create a major issue for chromosome segregation during cell division, which could cause DNA damage and result in cell death or cancer.
The researchers found that twisting a single chromatin fiber is much easier than twisting a double fiber. This means that the extra twist will preferentially go to the front, thus minimizing the intertwining of the two daughter DNA molecules.
“Although chromatin is normally considered an obstacle to replication,” Wang said, “our results show that chromatin also simplifies replication topology and thus facilitates replication dynamics. We feel that this is rather remarkable.”
In a separate experiment, the researchers found that an enzyme that untangles DNA (topoisomerase II) strongly prefers the single chromatin fiber in the front. The chromatin mechanics and topoisomerase activity seem to coordinate in a synergistic fashion to reduce daughter strand intertwining.
In order to understand how chromatin behaves mechanically, the researchers had to create new ways of handling it. Creating a substrate of braided chromatin fibers had not been previously attempted, because of the complexity of the task. Wang and her team used the angular optical trap tool that her group previously developed, as well as other methods to both create and work with both single and braided chromatin fiber substrates, enabling them to examine their torsional mechanical properties.
Paper co-authors include postdoctoral associates Tung Le, Xiang Gao and Jessica Killian from the Laboratory of Atomic and Solid State Physics; research specialist James Inman; graduate students Seong ha Park, Jaeyoon Lee and Ryan Badman; and Joyce Lee and James Berger of Johns Hopkins University School of Medicine.
Quartz cylinder fabrication was performed at the Cornell NanoScale Science and Technology Facility, a member of the National Nanotechnology Coordinated Infrastructure, which is supported by the National Science Foundation.
Linda B. Glaser is news and media relations manager for the College of Arts and Sciences.
Media Contact
Get Cornell news delivered right to your inbox.
Subscribe