Bacterial enzyme structure opens door for new antibiotics
By Krishna Ramanujan
Bacteria are growing increasingly antibiotic-resistant, but new research reveals how certain enzymes could be exploited to develop new classes of drugs to fight bacterial infections.
The study, “Structural Basis of Peptidoglycan Endopeptidase Regulation,” was published May 11 in Proceedings of the National Academy of Sciences.
The research examines how a type of enzyme – called endopeptidases – are regulated to break down cell walls in a process that allows bacteria to grow. The findings open the door to developing small molecules that exploit these enzymes to destroy bacterial cell walls and kill the pathogens.
“We need new approaches to find new antibiotics and the cell wall has been shown to be a very powerful target,” said Tobias Dörr, assistant professor in the Weill Institute of Cell and Molecular Biology and the Department of Microbiology and the paper’s senior author.
Bacterial cell walls are thin like eggshells but under constant pressure from within; they must also be pliable and must also allow the cell to grow, Dörr said. Endopeptidases allow the walls to grow by cutting holes in the cell walls, which are then filled by other proteins.
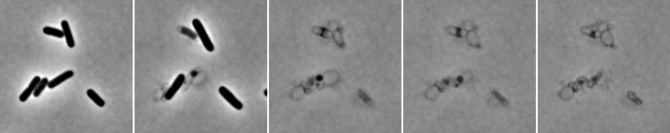
The study investigated how this system is regulated in cholera bacteria. Working with study co-authors Yuxin Mao, an associate professor in the Weill Institute, and Mark Saper, a biological chemist at the University of Michigan, the team used X-ray crystallography to find the structure of the endopeptidases. They identified two structures: an active open structure that allowed the enzyme to bind with the cell wall and another closed, inactive form. They then created mutations that promoted the open form, which they found enhanced activity. In collaboration with study co-author Felipe Cava at Umea University in Sweden, they found that remarkably, in this state, the enzymes destroyed cell walls in minutes.
This knowledge provides direction for designing a small molecule that prevents the active, open endopeptidase structure from switching back into an inactive form.
“We’re hoping that we can actually identify some chemical scaffolds that would allow us to lock that structure into the open conformation,” Dörr said.
Another option, he said, could be to find a way to keep the endopeptidases in its inactive form. “When inactive, the cell can’t grow and is trapped in its eggshell,” Dörr said. Such strategies could lead to new kinds of broad-spectrum antibiotics.
Penicillin and related antibiotics work by inhibiting cell wall synthesis – the patching up of the holes made by endopeptidases.
“If we can understand how endopeptidases and other enzymes make these holes, we might find other ways of inducing the same response,” Dörr said.
Jung-Ho Shin, a research associate in the Dörr lab, and Alan Sulpizio, a doctoral student in Mao’s lab, and Aaron Kelley, a former undergraduate in Saper’s lab, are co-lead authors.
The study is funded by National Institutes of Health.
Media Contact
Get Cornell news delivered right to your inbox.
Subscribe