Bacteria tails implicated in gut inflammation
By Krishna Ramanujan
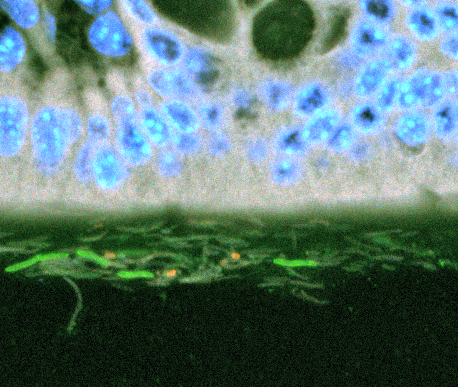
In healthy individuals, the only thing that separates the lining of the human gut from the some 100 trillion bacterial cells in the gastrointestinal tract is a layer of mucous.
But when gut bacteria do come in contact with cells on the gut’s surface, inflammation occurs, triggered by bacterial cell proteins. One particularly potent protein comes from the corkscrew-like tails of bacteria that propel the microorganisms. Though some of the most abundant bacteria in the gut use these tails (flagella) for propulsion, healthy guts have very low levels of the inflammation-causing proteins (flagellin). This suggests that some control mechanism must be in place to limit their production.
Now, a Cornell-led study published Nov. 13 in the journal Cell Host and Microbe reveals the complex dynamic between gut bacteria and the immune system that does, indeed, keep flagellin proteins in check.
The study was funded by the National Institutes of Health, Hartwell Foundation, Arnold and Mabel Beckman Foundation, and the Lucille and David Packard Foundation.
Read here to learn about how students in an art class used Ley's research to inspire their art.
“The gut pumps out six to 10 grams of antibodies a day directed against bacteria,” said Ruth Ley, assistant professor of microbiology and a senior author of the study. Tyler Cullender, a graduate student in Ley’s lab, is the paper’s first author.
Antibodies, which are produced by the immune system, attach to foreign substances. In the gut, antibodies called IgA are specifically directed against flagellin. The antibodies act like glue, creating a mucous layer that separates the bacteria from the gut, said Ley.
But, “the bacteria that get glued are not dumb,” said Ley. “Once they sense the antibodies – or that they are trapped – they stop making flagella and this probably limits how far they can swim” in the gooey layer, said Ley. With less flagellin proteins coming in contact with the gut lining, there is less inflammation.
The researchers studied mice that lacked a receptor called TLR5 for detecting flagellin proteins. Without the detector, these mice failed to produce adequate amounts of the IgA antibodies in the presence of flagellin. And without antibodies, many of the gut bacteria developed flagella and swam up against the cells of the gut.
When Ley and colleagues removed gut bacteria from these mice and added antibodies in the lab, the bacteria stopped producing flagella. “It worked to immobilize them,” Ley said.
The researchers’ next step is to confirm that the proximity of flagellin to the gut lining creates inflammation that triggers metabolic disease. Indeed, mice without the TLR5 receptor develop metabolic syndrome, a disease with a set of conditions, including increased blood pressure, high blood sugar, excess waist body fat and abnormal cholesterol levels, that occur together and increase the risk of heart disease, stroke and diabetes. Ley and colleagues had previously shown that the gut microbiota are required for the mice to develop these symptoms, and further research is necessary to show that the bacterial swimming is key to driving the disease.
Co-authors included researchers from Georgia State University, the University of Nebraska and Johns Hopkins University.
Media Contact
Get Cornell news delivered right to your inbox.
Subscribe