Details of HIV-1 structure open door for potential therapies
By Krishna Ramanujan
New research provides details of how the structure of the HIV-1 virus is assembled, findings that offer potential new targets for treatment.
The study, authored by a multi-institutional team led by Cornell researchers, was published Aug. 1 in the journal Nature.
The study reports that a small molecule called IP6, found in mammalian cells, plays key roles in the immature development of the virus (which occurs within an infected cell) and in the mature development of the virus (which occurs after the virus buds out of the cell membrane and cleaves from the cell).
“This small molecule acts in two different assembly steps in the pathway,” said Robert Dick, a postdoctoral researcher and the paper’s first author. Dick works in the lab of Volker Vogt, professor in the Department of Molecular Biology and Genetics and the paper’s senior author.
In both the immature and mature phases of the virus’ development, IP6 plays key roles in the pathways to create structural lattices. It facilitates assembly of an immature lattice while the virus develops inside the cell. This lattice breaks down after the virus buds and is cleaved from the cell membrane. At that point, IP6 also promotes the assembly of a second, mature lattice within the virus particle.
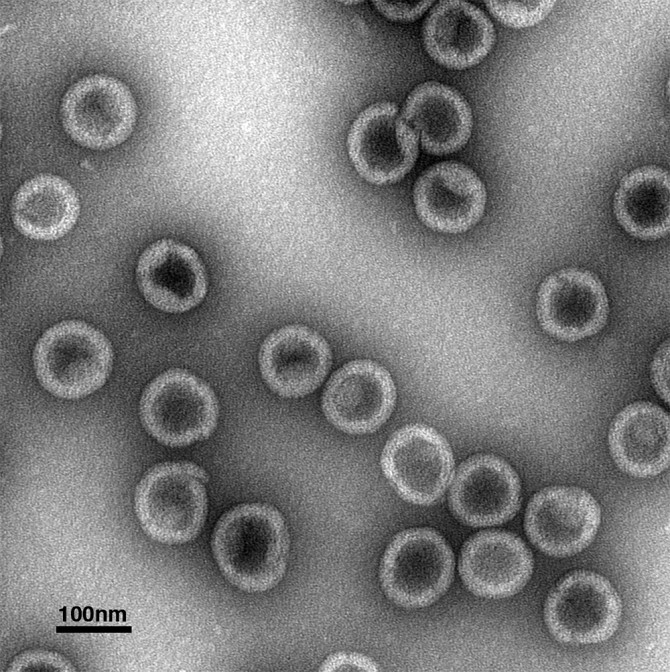
In the study, the researchers purified a structural protein, called a Gag protein, mixed it with a form of nucleic acid that acted as a template, and formed a protein virus lattice of the type found in the immature phase. Using a stain and an electron microscope, Dick and colleagues looked at the round virus particles (essentially just the protein virus lattice) on a grid.
The researchers repeated assembly reactions with the Gag protein and nucleic acid and a buffer in the presence and absence of IP6, Dick said: “Those first few experiments told us we were observing something pretty dramatic. The presence of IP6 greatly increased the number of virus particles we could detect.”
Still, the researchers were unsure where IP6 was acting on the Gag protein to have an effect, so they made different versions of the Gag protein and narrowed down the site of activity to make a prediction of where the small IP6 molecule was acting to create the lattice structure.
At that point, they approached co-authors Owen Pornillos and Barbie Ganser-Pornillos, both in the Department of Molecular Physiology and Biological Physics at the University of Virginia, who generated crystallographic data of the IP6 molecule interacting with the Gag structure.
In the mature phase, the researchers found that IP6 interacts at a site that is revealed after the virus particle cuts away from the cell. At that point, an entirely new lattice develops, made of different proteins than the immature lattice.
The findings open the door to possible new therapies. One option is for researchers to develop or identify compounds that are similar to IP6 and could bind to the same sites as IP6, thereby blocking it and preventing the virus from maturing.
“A cell can make millions of virus particles, but if they don’t go through the maturation process, they are not infectious,” Dick said.
Co-authors included teams of researchers from the University of Delaware; the European Molecular Biology Laboratory in Heidelberg, Germany; the Institute of Science and Technology in Klosterneuburg, Austria; and the University of Missouri.
This work was supported by the National Institutes of Health, the National Science Foundation and the German Research Foundation.
Media Contact
Get Cornell news delivered right to your inbox.
Subscribe