Cornell researchers reveal molecular basis of vision
By Krishna Ramanujan
Cornell and Stanford University researchers have solved the three-dimensional structure of a protein complex involved in vertebrate vision at atomic resolution, a finding that has broad implications for our understanding of biological signaling processes and the design of over a third of the drugs on the market today.
The researchers’ study, “Structures of the Rhodopsin-Transducin Complex: Insights into G-Protein Activation,” was published Aug. 22 in the journal Molecular Cell.
The findings illuminate how signals from photons (particles of light) get amplified in the eye. More importantly, the study provides insights into how the largest family of cell membrane proteins – G-protein-coupled receptors (GPCRs) – work in humans.
“They’re involved in almost all the biological processes in a human body – how we perceive light, taste, smell, or how the heart rate is regulated or muscles contract – and they are targets for over 30% of the drugs that are used today,” said Yang Gao, co-first author of the paper and a postdoctoral researcher in the lab of Richard Cerione, the Goldwin Smith Professor of Chemistry and Chemical Biology and co-senior author.
There are over 800 GPCRs in humans that signal through about 20 different G proteins. GPCRs are responsible for sensing a wide range of outside signals – such as hormones, light, and sense of smell and taste – and inducing corresponding responses inside the cell. In vertebrate vision, the GPCR rhodopsin is capable of detecting the signal from just one photon and through the activation of the G protein transducin and downstream effectors, amplify it 100,000 times.
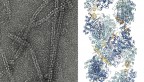
The researchers used cryo-electron microscopy to obtain atomic-resolution structures of the rhodopsin-transducin complex. The structures not only provide the molecular basis of vertebrate vision, but also reveal a previously unknown mechanism of how GPCRs in general activate G proteins.
“What we’ve learnt from these structures at an atomic level may be broadly applicable to other GPCR signaling systems,” said co-first author Sekar Ramachandran, a senior research associate in Cerione’s lab.
By learning more about how different receptors specifically couple with different G proteins, the researchers hope to gain insights into designing drugs that specifically regulate GPCR signaling. A lot of drug side effects occur when therapies are not specific enough and target both harmful and beneficial pathways, Yang said.
Hongli Hu, a postdoctoral researcher in Stanford’s Department of Structural Biology, is a co-first author; Georgios Skiniotis, professor of molecular and cellular physiology and of structural biology at Stanford, is a co-senior author.
The study, which was featured as Editor’s Choice in the Sept. 3 issue of the journal Science Signaling, was funded by the National Institutes of Health.
Media Contact
Get Cornell news delivered right to your inbox.
Subscribe