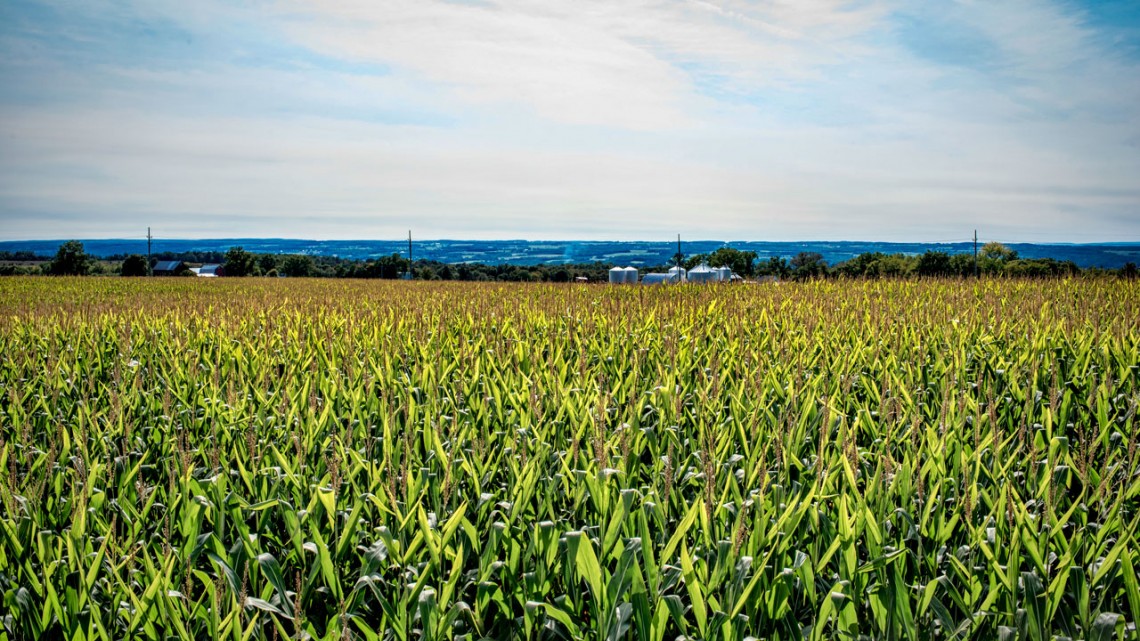
Corn grows at Musgrave Farm Research station in Aurora, New York.
New genomic tools help improve staple crops worldwide
By David H. Freedman
Plant scientists have made tremendous advances over the past decade in identifying genomic data that can drive crop-breeding decisions. But while those genomics tools have become commonplace among breeders in highly industrialized countries, they haven’t been widely accessible in developing nations.
To boost the availability of plant-breeding tools, a Cornell-based global program – the Genomic Open-source Breeding informatics initiative, or GOBii – has been developing better breeding tools and expanding access to genomic databases.
The initiative is a collaboration between Cornell, researchers at the Boyce Thompson Institute, and researchers and scientists from around the world. The program launched in 2015 with an $18.5 million grant from the Bill & Melinda Gates Foundation and will operate until October of this year.
GOBii’s goal is to speed up the breeding process for crops that are important in developing regions, in addition to improving yield, nutritional quality, disease resistance, weather resistance and other desirable traits. The program’s new software and databases are starting to be deployed around the world, and early breeding efforts have indicated that they can slice at least a year off what can take as many as five years.
The program, which initially focused research efforts on ways to support India, Mexico and the Philippines, has recently expanded to countries in Africa. The new databases include information on crops such as chickpeas, maize, millets, rice sorghum and wheat.
“A lot of the crop varieties used in South Asia and Africa were developed two or more decades ago,” said Kelly Robbins, assistant professor of plant breeding and genetics in the College of Agriculture and Life Sciences. “Replacing these older varieties with new and improved varieties will have a significant impact on the quality of life of people living in these regions.”
Genomics tools work by combing through databases of genetic data to identify particular “molecular markers,” or small sections of a plant’s DNA, that influence specific traits. A breeder with the right tools and data can examine seeds or seedlings for desirable molecular markers, and then predict whether the full-grown plant will have the ideal characteristics – rather than waiting for observational results at the end of the growing season.
“Markers can be very predictive of traits, so you can make breeding decisions much earlier in the breeding cycle and with more accuracy,” said Liz Jones, director of GOBii.
After the grant ends, GOBii’s work will be integrated into an even bigger project, Excellence in Breeding. Run by CGIAR, a global nonprofit for agricultural innovation, Excellence in Breeding aims to train people on the ground in key countries to ensure that the tools are utilized in ways that improve farming and local nutrition.
Robbins will also be working with Excellence in Breeding. One of the researchers’ ongoing challenges is to better understand which traits local farmers find desirable in specific crops – especially since those needs often vary widely from community to community.
“GOBii made great progress in developing the breeding capabilities,” Robbins said. “But getting to routine adoption in those regions is going to be the new focus going forward.”
For a longer version of this story, visit the College of Agriculture and Life Sciences website.
David H. Freedman is a freelance writer for the College of Agriculture and Life Sciences.
Media Contact
Get Cornell news delivered right to your inbox.
Subscribe