Spatial maps give new view of gut microbiome
By David Nutt
What microbes are in your gut, and where?
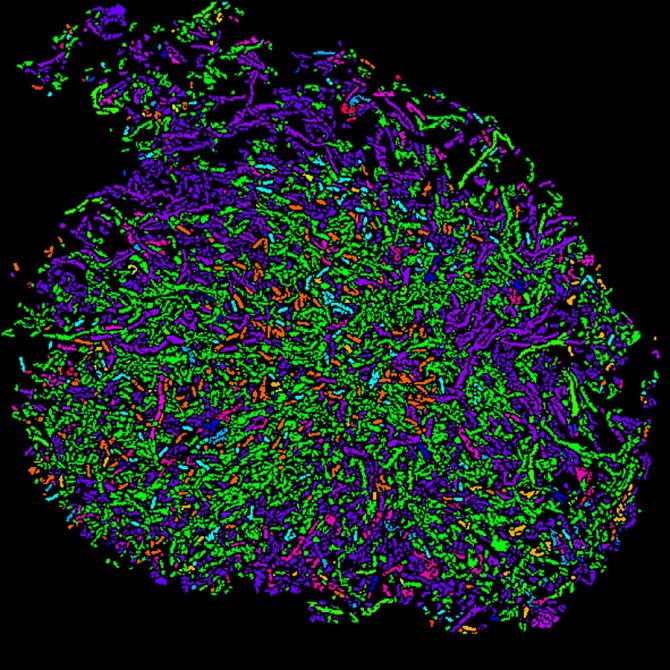
Cornell researchers developed an imaging tool to create intricate spatial maps of the locations and identities of hundreds of different microbial species, such as those that make up the gut microbiome. The tool will help scientists understand how complex communities of microorganisms interact with each other and also their environment, which is to say, us.
The team’s paper, “Highly Multiplexed Spatial Mapping of Microbial Communities,” published Dec. 2 in Nature. The paper’s lead author is doctoral student Hao Shi, M.Eng. ’18.
“There are communities of bacteria that live in our bodies and play an important role in human health and biology, and there’s a rich diversity of these microbes. We know this from technologies such as DNA sequencing that create lists of the bacterial species that are present in a community,” said Iwijn De Vlaminck, the Robert N. Noyce Assistant Professor in Life Science and Technology in the Meinig School of Biomedical Engineering, and the paper’s senior author.
“However, there are very limited tools to understand the spatial interactions between these microbes, and those are quite clearly important to understand the metabolism of these communities, and also how these microbes interact with their host,” he said.
De Vlaminck and Shi set out to create their imaging method by using a two-step process called high phylogenetic resolution microbiome mapping by fluorescence in situ hybridization (HiPR-FISH). They collaborated with the labs of co-authors Warren Zipfel, associate professor of biomedical engineering, and Ilana Brito, assistant professor and the Mong Family Sesquicentennial Faculty Fellow in Biomedical Engineering, to incorporate additional imaging and microbiome expertise.
To locate the microbial communities, the researchers designed oligonucleotide probes that target specific bacteria cells based on the presence of a signature gene sequence, 16S ribosomal RNA, and they made another group of probes that label the cells with fluorophores. Then the team used confocal microscopy to light up the fluorescent markers with lasers, and they used machine learning and custom software to decode the fluorescence spectra and interpret the images, resulting in an efficient and cost-effective technology with single-cell resolution.
The researchers created the palette for their spatial maps with a mixture of 10 basic colors that could “paint” a total of 1,023 possible color combinations of E. coli, each fluorescently labeled with a unique binary barcode.
“The imaging itself leads to very beautiful, rich images with all bacterial cells in different colors,” De Vlaminck said. “But to allow the quantitative understanding of microbe interactions, the distances between cells, cluster sizes and so on, you need to be able to interpret these in an automated way by a computer so that you can convert this image into a digitized representation of the community.”
The team applied their technology to two different systems: the gut microbiome in mice and the human oral plaque microbiome. In the case of the gut microbiome, they were able to demonstrate how the spatial associations between different bacteria are disrupted by antibiotic treatment.
Spatial mapping could be an important tool for studying and possibly treating a range of diseases in which bacteria are a major culprit, such as inflammatory bowel disease, colorectal cancer and infection.
“We’d like to dig deeper into the biology of systems where microbiomes play important roles and try to understand how these kinds of spatial dynamics change when you have a disease in progression,” Shi said. “We want to see if that offers any clues and therapeutic insights that we can harness to help people.”
Co-authors are doctoral student Benjamin Grodner; research associate Joan Sesing Lenz; and research support specialist Qiaojuan Shi.
The research was supported by the Kavli Institute at Cornell for Nanoscale Science and the National Institutes of Health (NIH). The researchers made use of the Cornell BRC-Imaging Facility, which is supported by New York State Stem Cell Science and the NIH.
Media Contact
Get Cornell news delivered right to your inbox.
Subscribe